The LCGC Blog: Multipath Liquid Chromatography–Mass Spectrometry: A Veritable Pandora’s Box
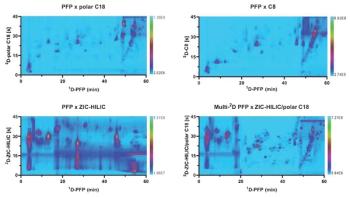
For several years, our group has been working on a concept that we have termed multipath liquid chromatography (LC). The main idea is to target multiple classes of compounds following a single injection of a sample, the components of which are segregated on-line and directed to separate appropriate paths for simultaneous separation; the streams are then recombined for detection. I believe that this approach would be powerful for biomarker quantitation, where it would be more informative to track both metabolite and protein biomarkers to better define a disease state, or in the case of antibody–drug conjugate (ADC) development, where the metabolism of the ADC might involve understanding both the levels of the released drug and the remaining protein.
I suppose that Pandora’s Box usually has a negative connotation, but in this case it is not so. For several years, our group has been working on a concept that we have termed multipath liquid chromatography (LC). The main idea is to target multiple classes of compounds following a single injection of a sample, the components of which are segregated on-line and directed to separate appropriate paths for simultaneous separation; the streams are then recombined for detection. I believe that this approach would be powerful for biomarker quantitation, where it would be more informative to track both metabolite and protein biomarkers to better define a disease state, or in the case of antibody–drug conjugate (ADC) development, where the metabolism of the ADC might involve understanding both the levels of the released drug and the remaining protein. Performing these analyses with one injection would certainly be preferable to having separate methods (and likely, sample preparations) to target each class separately.
The initial embodiment of the multipath system uses restricted-access media (RAM) to separate proteins from small molecules; each class is then directed to its own LC separation and the separations are then joined back together via a tee connection to enter the mass spectrometer detector (1). There is a lot of room to make this concept more complex and more powerful.
The reason this approach has taken so long to develop is that the overarching concept contains many little intricacies, each of which needs to be studied and optimized. We would like to be able to implement automated method scouting to reduce method development time; to target and quantify intact proteins, to avoid extra steps and uncertainties associated with protein digestion; and to incorporate comprehensive multidimensional LC (LCxLC) in each of the paths. We still have a lot of work to do to be able to reach these goals, but we have made headway in many aspects.
I have written previously about our efforts to use automated method scouting to streamline method development (2). Using a diverse set of small molecules, in two separate studies we screened a series of commercial columns to characterize their performance in reversed-phase mode (3,4). In the second case, we also screened a column set in aqueous normal-phase mode. The take-home message was that automated method scouting, using a generic mobile phase gradient to determine which column provides the best selectivity and peak shapes for separating a set of desired analytes, is a much more efficient route to method development than sticking with one column and trying to find a mobile phase that works. After all, changing the stationary phase is the most effective way to affect selectivity for small-molecule separations.
A bit more challenging was figuring out how to target intact proteins using triple-quadrupole mass spectrometry (MS). I also talked about this before in another LCGC Blog installment (5). It was a good thing that we started working on this approach before we really consulted people. In the literature, there was very little to no precedent for using multiple reaction monitoring in triple-quadrupole MS, like what you would do for small molecules, on intact proteins. Most people claimed it would not work, but we were successful.
We were able to create a reasonably sensitive, but highly specific, means for top-down quantitation of intact proteins (6). The most important part of the experimental design turned out to be modulating the collision energy to a point where not all of the precursor ion was fragmented; under those conditions, a series of highly reproducible and specific product ions could be generated. We have not analyzed a protein for which this approach fails to work.
We also spent considerable effort to better understand ion transmission properties of proteins versus small molecules in the gas phase, inside the triple-quadrupole analyzer (7,8). Overall, we concluded that there probably is some opportunity to develop a system that is tailored and more amenable to protein analysis, since essentially all triple-quadrupole instruments are dedicated to (and presumably tested for) small-molecule analysis.
Figuring out how to detect the intact proteins allowed us to return back to studying their separation. A dearth of literature exists on the topic, but much less of that previous work considered direct detection by MS. Thus, many reported intact protein separation conditions from the 1980s and 1990s incorporated non-MS-compatible mobile-phase conditions. We were able to study both stationary-phase and mobile-phase effects on the separation of a model set of proteins and to define some generally good conditions for MS-compatible separations (9). The most important aspect of this work was to realize that a compromise needed to be made with mobile-phase additives. Formic acid is good for MS sensitivity to detect the intact proteins, but it generally provides for poor peak shapes. Trifluoroacetic acid is a common additive for generating good peak shapes for proteins, but it can suppress electrospray ionization response. We found that a combination of the two, where just a small amount (0.05%) of trifluoroacetic acid is used along with a standard amount (0.1 or 0.5%) of formic acid, to provide excellent performance.
We continue down the path. Just this week at the HPLC conference in Washington D.C., I presented some of our recent headway toward generating separation conditions orthogonal to those I discussed above. The impetus for those efforts was to move toward LCxLC of intact proteins, and I was also able to show some good progress on that front. We are currently preparing manuscripts to more comprehensively report on those advancements. Stay tuned for more development on the multipath LC concept.
References
1. D.K. Appulage, E.H. Wang, B.J. Figard, and K.A. Schug, J. Sep. Sci.41, 2702–2709 (2018).
2. K.A. Schug, The LCGC Blog, August 7, 2017.
3. D.K. Appulage, E.H. Wang, F. Carroll, and K.A. Schug, J. Sep. Sci.39, 1638–1647 (2016).
4. D.K. Appulage and K.A. Schug, J. Chromatogr. A1507, 115–123 (2017).
5. K.A. Schug, The LCGC Blog, Aug. 9, 2016.
6. E.H. Wang, P.C. Combe, and K.A. Schug, J. Am. Soc. Mass Spectrom.27, 886–896 (2016).
7. K.A. Schug, The LCGC Blog, June 28, 2017.
8. E.H. Wang, D.K. Appulage, E.A. McAllister, and K.A. Schug, J. Am. Soc. Mass Spectrom.28, 1977–1986 (2017).
9. E.H. Wang, Y. Nagarajan, F. Carroll, and K.A. Schug, J. Sep. Sci.39, 3716–3727 (2016).
Kevin A. Schug is a Full Professor and Shimadzu Distinguished Professor of Analytical Chemistry in the
Department of Chemistry & Biochemistry at The University of Texas (UT) at Arlington. He joined the faculty at UT Arlington in 2005 after completing a Ph.D. in Chemistry at Virginia Tech under the direction of Prof. Harold M. McNair and a post-doctoral fellowship at the University of Vienna under Prof. Wolfgang Lindner. Research in the Schug group spans fundamental and applied areas of separation science and mass spectrometry. Schug was named the LCGC Emerging Leader in Chromatography in 2009 and the 2012 American Chemical Society Division of Analytical Chemistry Young Investigator in Separation Science. He is a fellow of both the U.T. Arlington and U.T. System-Wide Academies of Distinguished Teachers.
Newsletter
Join the global community of analytical scientists who trust LCGC for insights on the latest techniques, trends, and expert solutions in chromatography.