- The Column-08-13-2018
- Volume 14
- Issue 8
Tips & Tricks GPC/SEC: UV–vis Detection
The most commonly applied detector in gel permeation chromatography/size-exclusion chromatography (GPC/SEC) is the differential refractive index detector, RI. How UV–vis detection, if applicable, adds true value to GPC/SEC applications is discussed in this instalment of Tips & Tricks.
The most commonly applied detector in gel permeation chromatography/size-exclusion chromatography (GPC/SEC) is the differential refractive index detector, RI. How UV–vis detection, if applicable, adds true value to GPC/SEC applications is discussed in this instalment of Tips & Tricks.
Gel permeation chromatography/sizeâexclusion chromatography (GPC/SEC) is usually used to determine the molar mass distribution (MMD) and molar mass averages of a polymer sample of known chemistry. Peak identification and concentration quantification for each separated peak, the most common tasks in high performance liquid chromatography (HPLC), are often of less interest. To determine the MMD, GPC/SEC requires detectors that measure the relative or-when using light scattering or viscometry detection-absolute concentration in every chromatographic slice.
Refractive index detectors, RIs, are the most common GPC/SEC detectors because of their universal applicability; they do not require chromophores to be present in the analyte (1). However, for UV–vis active samples, in general, for analytes containing unsaturated bonds, aromatic groups, or functional groups with heteroatoms, the use of a UV–vis detector offers several advantages.
UV–vis detectors are thus used for both natural (peptides, proteins) and synthetic macromolecules (for comparison see table 1 in reference 2). In addition, the combination of UV–vis and RI detection is widely applied because it offers additional information, for example, about the variation of composition with elution volume.
Types of UV–vis Detectors, Applications, and Advantages
Typically, UV–vis detectors work with light from a deuterium (UV) or tungsten (visible) lamp. The Beer-Lambert law describes the relation between the absorbance (A) and the analyte concentration (c):
A = ε · c · l [1]
ε is the molar absorption coefficient (dm3 mol-1 cm-1) and l is the path length of the light through the flow cell (cm). If ε is not known, solutions of known concentration can be used for its determination and for calibrating the detector response (which is required when working with light scattering detectors).
As mentioned above, the most important task in GPC/SEC is determination of the slice concentration. Detection at one specific wavelength is often fully sufficient and the majority of applications can therefore be run with single wavelength UV–vis detectors. Example applications are gelatin–hydrolyzed collagen, which can be easily characterized using a single wavelength detector operated at 214 nm, or synthetic polymers bearing aromatic moieties, which can be detected at 254 nm.
Dual wavelength detection can be applied in protein analysis to additionally quantitate the amount of protein in the solution. The peptide bonds absorb at around 205 nm. Detection at 214 nm and 280 nm allows more detail to be obtained as UV–vis absorbance between different proteins varies strongly at 280 nm and is related to the actual content of tyrosine, tryptophan, and cysteine.
Photodiode array detectors (PDAs or DADs), which can scan a selectable range of wavelengths, are quite common in HPLC. However, there is only a limited number of applications in GPC/SEC where this functionality is required. In those applications spectra are used to identify oligomeric species, for example, in wood resins or polymer additives. Additional concentration detectors (RIs) are often used to obtain the molar mass distribution results simultaneously to the spectral information. Sometimes the PDA/DAD scan functionality is used to determine a wavelength that can later be used for the specific detection.
However, in all cases of UV–vis detection, users should verify upfront that all analyte components can be detected. UV–vis detection alone has a high risk of undetected components because many macromolecules do not exhibit any chromophores. The verification can be done with the additional use of an RI detector.
The inset in Figure 1 shows 3D spectra taken from 240 nm to 390 nm for a GPC separation in tetrahydrofuran (THF). The UV trace (red) at 254 nm detects those components bearing suitable chromophores. The component eluting at 22.5 mL shows an additional absorbance around 300 nm, indicating a chemically different entity. The last component at 23.2 mL is not seen by DAD/PDA because of the absence of suitable chromophores; however, the component is detected by the RI (green trace).
One advantage of UV–vis detectors when compared to other GPC/SEC detectors is that they have a small cell volume. The lower band broadening of smaller cells can be beneficial if the resolution of close peak pairs is of concern. The UV–vis detector can (and should) be placed as a first detector in a detector train because of its lower contribution to band broadening. Its cell is also very pressure stable and the signal does not depend on back pressure (3).
UV–vis detectors have a high linearity, selectivity, and sensitivity and are suitable if scientists have verified upfront that all analyte components can be detected.
Detector Combinations with UV–vis Detectors
UV–vis detectors have a high selectivity. Appropriate setting of the wavelength may allow selective detection, for example, of only one type of comonomer in a copolymer or of a specific end group. This makes UV–vis detectors a very valuable addition in detector combinations. The protein application mentioned above is an example of UV–vis–UV–vis dual detection; more generally applicable is the combination of a UV–vis detector with an RI detector
UV–vis–RI combinations are often used in copolymer analysis to determine the chemical composition distribution in a given copolymer (4). Figure 2 shows an example of a chromatogram of a graft copolymer consisting of polymethyl methacrylate (PMMA) and polystyrene (PS) measured in THF. The universal RI detector responds to both monomer units, MMA and styrene. However, the UV–vis detector set to a wavelength of 254 nm selectively detects the styrene repetition units, which bear the chromophore. The detector traces appear to be shifted relative to each other although the inter-detector delay between UV–vis and RI has been properly accounted for. This shift of detector traces is a consequence of the variation of the chemical composition with elution volume and the different responses of both detectors.
This difference in response to the same analyte allows more to be learned about copolymers. The simple fact that the (normalized) detector traces of both concentration detectors do not superimpose is an indication of chains of different chemical composition. At low elution volumes, the (normalized) RI signal is higher than the UV signal, while the opposite is true at elution volumes exceeding 8.5 mL. To a first approximation the RI signal is proportional to the total mass concentration of molecules eluting from the column. Thus, at a given concentration the species eluting at low elution volumes bear less absorbing groups than at high elution volumes. In other words: the larger molecules (low elution volumes) are richer in MMA, while the shorter chains (high elution volumes) are rich in polystyrene, as indicated by the higher UV absorbance.
While the above explanation is qualitative only, quantitative information can be obtained as well. This opens up new calibration possibilities for block copolymers. Based on the two detector signals, the chemical composition can be calculated for each elution volume, allowing a copolymer calibration for the specific composition from the calibration curves of the two homopolymers to be established, which finally permits molar mass determination for copolymers (5).
Another option in UV–RI detection is the selective detection of end groups by choosing a suitable UV–vis wavelength. This approach is applied in the European Pharmacopoeia for the molar mass determination of low-molecular-weight heparins. Figure 3 shows an example chromatogram of an end group modified heparin reference material. The end group of this calibration standard can be detected by 234 nm. A UV–vis detector set to this wavelength shows a signal that is related to the number of chains eluting from the column. The RI detector-used in addition-detects every repetition unit of the heparin chains, so that the signal is related to the mass concentration of the chains passing the detector. The fact that the UV signal is proportional to the number concentration, while the RI signal is proportional to the mass concentration of eluting chains is the reason for the strongly differing detector traces. As the number average molar mass (Mn) for this standard is known, it is now possible to create a heparin calibration curve based on the UV–RI ratio. The obtained calibration curve can subsequently be applied for molar mass determination of heparins using the RI signal only.
This end group calibration method can be applied to all samples where it is possible to specifically detect end groups and where every chain definitely bears one end group. The only additional information required then is the Mn.
On the other hand, it is also possible to measure the end group distribution within a sample with UV–vis-detectable end groups when using UV–vis–RI dual detection.
Summary
- UV–vis detectors are linear, sensitive,
- and selective detectors. They are ideal GPC/SEC detectors if all analyte components have chromophores.
- Single wavelength UV–vis detectors are often sufficient. PDA/DAD areonly used for a limited number of applications but provide UV–vis spectra for all single components in a run.
- Selecting a specific wavelength to detect one comonomer in a copolymer or an end group adds additional value when using dual detection with RI.
- UV–vis detectors can be set to detect the repetition unit (weight average signal) or to detect end groups (number average signal). The latter offers opportunities for calibration to obtain true masses.
References
- D. Held, The Column13(17), 24–27 (2017).
- D. Held, J. Preis, and F. Gores, The Column 11(22), 20–23 (2015).
- D. Held and P. Kilz, The Column8(18) 9–12 (2012).
- R. Leinweber, P. Montag, J. Preis, and W. Radke, Journal of Chromatography A1473, 76–82 (2016).
- P. Kilz, The Column 6(10), 12–17 (2010).
Daniela Held studied polymer chemistry in Mainz, Germany, and works in the PSS software and instrument department. She is also responsible for education and customer training.
E-mail:
Articles in this issue
over 7 years ago
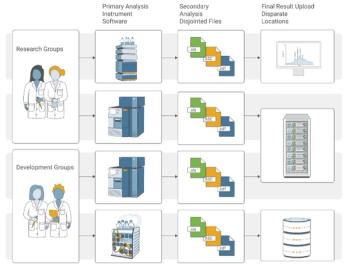
Analytical Life Cycle Management— The Coming Revolutionover 7 years ago
Optimizing Splitless GC Injectionsover 7 years ago
Advancing Chromatography Methods for Cannabis Analysisover 7 years ago
Forensic Profiling of Human Odour Using GC×GC–MSover 7 years ago
Sciex Donates to World Cancer Research Fundover 7 years ago
Biodiesel-Diesel Blend Analysis Using UFGCover 7 years ago
Vol 14 No 8 The Column August 2018 North American PDFover 7 years ago
Vol 14 No 8 The Column August 2018 Europe and Asia PDFNewsletter
Join the global community of analytical scientists who trust LCGC for insights on the latest techniques, trends, and expert solutions in chromatography.