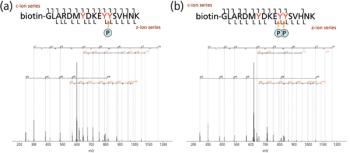
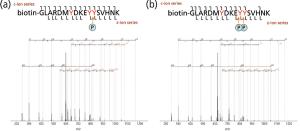
Protein and peptide analysis via tandem mass spectrometry (MS-MS) has resulted in a wealth of information regarding protein identification, structure, and abundance levels over the past 10 years. Techniques such as neutral loss scanning and collision-induced dissociation (CID) have been especially helpful in facilitating the identification of a multitude of previously unknown sites of protein phosphorylation. However, many of the techniques used to obtain this information are labor intensive and work inconsistently. To address this problem, much effort has been put forth to find alternative methods of fragmenting peptides and proteins that are less difficult and applicable to a wide gamut of peptide classes. Examples of recently developed dissociation techniques include infrared multiphoton dissociation (IRMPD) and electron transfer dissociation (ETD). The implementation of these new techniques has widened the spectrum of peptides amenable to tandem mass spectral analysis.