- May 2022
- Volume 18
- Issue 05
- Pages: 13–16
Accelerating the Understanding of Metabolomics in Disease Through Improved Sensitivity
Accelerating the understanding of metabolomics in disease is increasingly high on the agenda for many researchers looking to develop their drug discovery efforts into clinical trials. With personalized medicine the goal that many are striving to achieve, metabolomic technologies must continue to develop improved sensitivity if they are to make a real difference to the drug research of the future.
Metabolites are the chemical entities that are transformed during metabolism in the body, and they provide a functional readout of cellular biochemistry. Metabolites are the products of metabolic pathways and include small molecules such as lipids, sugars, nucleotides, and amino acids. With mass spectrometry (MS)-based metabolomics techniques, thousands of metabolites can be quantitatively measured from minimal amounts of biological material, which has subsequently enabled systems-level analyses. Global or untargeted metabolite profiling is revealing new discoveries linking cellular pathways to biological mechanisms, shaping the understanding of cell biology, physiology, and medicine.
Metabolomics plays a major role in clinical practice, as it represents more than 95% of the workload in clinical laboratories worldwide (1). Although relatively new compared with its genomic and proteomic predecessors, research into metabolomics has already led to the discovery of biomarkers for disease diagnosis, fundamental insights into cellular biochemistry, and observations related to disease pathogenesis.
The Metabolomic Challenge
One of the biggest challenges facing untargeted metabolomics research is reliable compound identification to ensure accurate interpretation of data. Due to the chemical similarity of metabolites (isobars and isomers), identification by MS or chromatography alone can often be difficult for researchers looking into the behaviour of metabolites. The challenge is that isomers can have the same mass, charge, and physical properties, but can vary widely in their bioactivity, making both separation and identification a time- and resource-intensive process.
Another challenge is that the metabolome is extremely sensitive, meaning a person’s metabolic phenotype is influenced by environmental factors. Data can also be more ambiguous because metabolites are the products of upstream biological transformations of molecules in a similar mass range, which in turn presents the need for higher sensitivity to obtain broad metabolite coverage and relative concentrations. The field of metabolomics is much newer than proteomics and researchers also experience significant challenges in the interpretation of data, for example, in the interpretation of fragmentation data.
Ion mobility spectrometry–mass spectrometry (IMS-MS) has emerged as a powerful technology to address such issues, and the use of IMS-MS in untargeted metabolomics has improved the separation of metabolite isomers and supports metabolite identification through generation of orthogonal data, such as collision cross section values.
New technology is continually being developed to realize the biological relevance of these metabolites, to delve deeper into their behaviours, and to look further into new metabolomic possibilities.
Metabolomics and the Study of Lipidomics
Alterations at the metabolome level reflect disturbances in preceding biological cascades, bridging the gap between the genome and phenotype. Changes at this level can lead to the onset of disease symptoms, making metabolomics an essential diagnostic and prognostic tool in investigating the mode of action of chemical compounds and obtaining an in-depth understanding of the impact of infection, for example (2). Current metabolomics research focuses on a range of challenging diseases, including Alzheimer’s disease and cancer.
Scientists are looking further into metabolomic research to examine how the cells in our body behave and what this could mean as we look towards a future of personalized medicine. The acceleration of this research has driven new developments in lipid biochemistry. Lipids are fatty small molecules that share common physical and chemical properties, and the abundance of lipids and different lipid classes is key to metabolic regulation. Lipids studied using metabolomic approaches—lipidomics—are an ideal subject for metabolomic measurements. Lipids also allow scientists to gain further insight into genomes and genetic polymorphisms (3).
As metabolomic possibilities progress, the study of lipidomics has become a prominent source for collecting and measuring data within the metabolome. Because the abundance of lipids in typical samples, such as plasma extracts, can vary considerably, analytical methods with high sensitivity and dynamic range are required. One novel method focuses on coupling a vacuum insulated probe heated ion source (VIP-HESI) with a mass spectrometer. This technology looks at enhancing the limit of detection (LOD) and the dynamic range, as well as examining the lipid identification annotation capability. It enables sensitivity gains of up to 16-fold by efficient desolvation, enabling new applications across all small molecules (4).
Looking further into lipids is crucial to further understanding challenging diseases. Lipids are involved in biological processes such as blood clotting, mediating inflammation, or as signalling molecules, which makes them an interesting target for the discovery of new biomarkers. Insights such as this are essential for developing new treatments and for personalized medicine (5). The discovery of new biomarkers increases the likelihood of finding real-life outcomes for patients as this data progresses into the clinic.
It is key that research progresses in the field of metabolomics, and, within that, lipidomics, to transform insights and cellular findings into new drugs. In turn, personalized medicine becomes a brighter prospect to combat a range
of diseases.
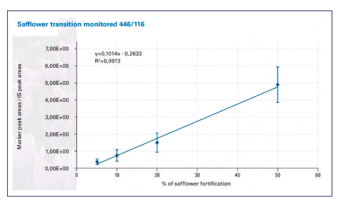
Data from an experiment using VIP-HESI is highlighted in Figure 1, showing the difference that it makes to the analysis and sensitivity of lipids across different sample concentrations (4). This technology allows data from lipids to be acquired through a deeper and broader analysis than in previous lipidomic workflows, with the figure showing a higher number of lipid identification annotations for untargeted workflows (4).
Looking Towards the Future of Personalized Medicine
As new technologies evolve and researchers look to optimize equipment to accelerate their studies, it is important that software and analytics keep pace to support the abundance of new data. Looking to a future in which personalized medicine is perceived to be within reach, advances in metabolomics research are significant in discovering new biomarkers to guide insight into our genetic make‑up. With the advancement of MS, as well as technologies such as nuclear magnetic resonance (NMR),
further strides can be made in clinical practice.
References
- M. Jacob, A.L. Lopata, M. Dasouki, and A.M. Abdel Rahman, Mass Spectrom. Rev.38(3), 221–238 (2019).
- V. Tounta, Y. Liu, A. Cheyne, and G. Larrouy‑Maumus, Molecular Omics 17(3), 376–393 (2021).
- J.B. German, L.A. Gillies, J.T. Smilowitz, A.M. Zivkovic, and S.M. Watkins, Current Opinion in Lipidology18(1), 66–71 (2007).
- M. Szesny, L. Woods, S. Meyer, A. Korf, N. Kessler, and A. Barsch, Improved sensitivity and higher lipid annotation ID capabilities using a new vacuum insulated heated ESI source (Bruker, 2021).
- P. Schmitz, Lipidomics (online). Available at:
https://www.uni-due.de/aac/lipidomics.php (Accessed 8 November 2021).
Lucy Woods is Business Unit Manager Phenomics and Metabolomics at Bruker Daltonics.
E-mail:
Website:
Articles in this issue
about 4 years ago
Rising Stars of Separation Science: Valentina D’Atriabout 4 years ago
Effectiveness of High-Resolution Ion Mobility for Complex Analysesabout 4 years ago
Improved Characterization of Malodours in Recycled Plasticsabout 4 years ago
ISSS 2022 and HPTLC 2022 Event Previewsabout 4 years ago
Thermo Fisher Scientific Continues Expansionabout 4 years ago
PharmaZell and Novasep Announce Merger Completionabout 4 years ago
Investigating Ciguatera Using LC–MS/MSabout 4 years ago
Analyzing Road-Associated Microplastics Using Py-GC–MS