- The Column-01-19-2016
- Volume 12
- Issue 1
The Power of 2D LC
Protein biopharmaceuticals have seen an enormous growth in the last decade, and as a result, separation scientists are giving increased attention to methods for characterizing biopharmaceuticals. One powerful technique for analyzing proteins is two-dimensional liquid chromatography (2D LC). Gerd Vanhoenacker of the Research Institute for Chromatography (RIC) in Kortrijk, Belgium, has been conducting research into peptide mapping of therapeutic monoclonal antibodies (mAbs) using 2D LC. He recently spoke to The Column about this work.
Protein biopharmaceuticals have seen an enormous growth in the last decade, and as a result, separation scientists are giving increased attention to methods for characterizing biopharmaceuticals. One powerful technique for analyzing proteins is two-dimensional liquid chromatography (2D LC). Gerd Vanhoenacker of the Research Institute for Chromatography (RIC) in Kortrijk, Belgium, has been conducting research into peptide mapping of therapeutic monoclonal antibodies (mAbs) using 2D LC. He recently spoke to The Column about this work.
Q. In a recent article you mention that within the current decade you expect a much larger share of drug approvals to be of a biological nature, specifically monoclonal antibodies.1 Why do you think this is the way the therapeutics market is going?
A: Protein biopharmaceuticals have emerged as important therapeutics for the treatment of cancer, cardiovascular diseases, diabetes, infection, and inflammatory and autoimmune disorders. Given their obvious benefits in terms of safety and efficacy, they are reshaping the pharmaceutical market. The majority of these proteins are, and will be, monoclonal antibodies (mAbs) and many of these have already received approval in Europe and the United States.
In a recent supplement from LCGC Europe, edited by Pat and Koen Sandra, a brief overview of current and future trends in drug development and sales was given.2 Trends for different classes of pharmaceuticals speak for themselves: Small-molecule drug sales are stagnating while recombinant protein pharmaceuticals sales have increased by over 25% between 2008 and 2013. Sales for mAbs have nearly doubled over the same period of time. Big pharma organizations are now more and more focused on biopharmaceutical products. Of course, they will continue to invest in the development of key small molecule formulations, but a major part of their research is now focused on large biomolecule drugs in general, and mAbs in particular.
Q. This shift would obviously represent an analytical challenge. Do you feel the pharmaceutical industry is sufficiently experienced and prepared for such a shift?
A: The analysis of these big, heterogeneous biomolecules will often require a shift in analytical approach and technique compared to the analysis of small-molecule pharmaceuticals. This shift in approach is needed at nearly all levels going from sample storage and preparation to analysis and data treatment. Most regulatory guidelines were originally developed for smallâmolecule drug products and formulations. To enable researchers in academia and industry to set up and validate their methodologies, separate guidelines for biopharmaceutical drug substances and products have been issued such as the International Conference on Harmonization (ICH) Q6B guideline on test procedures and acceptance criteria for biotechnological and biological products, and ICH Topic Q5C, D, and E.
I believe there are opportunities for products and techniques that can automate or accelerate all steps involved in the bioanalytical process. A lot of progress has been made over the last decades, but the present achievements are definitely not the endpoint. Advances towards faster analyses and even more accurate and detailed data generation will be necessary and will be made. Innovations in mass spectrometry (MS), chromatography (particularly ultrahighâpressure liquid chromatography [UHPLC] and twoâdimensional liquid chromatography [2D LC]), and software are most important. The first two might seem obvious, but the last is equally important. A critical part of biopharmaceutical analysis lies in the data handling and interpretation. Data analysis software is playing a very prominent role in research on various biopharmaceuticals and this is a major difference compared to the analysis of small-molecule pharmaceuticals. It is a field in which various instrument and software developers are investing considerably.
Q. Peptide mapping is obviously a very useful technique, can current one-dimensional (1D) LC methods deal with the large amount of peptides that comprise a large protein drug?
A: The power of state-of-the-art 1D LC techniques should not be underestimated. A well-developed (U)HPLC method can be very valuable and robust for peptide mapping. However, when sample complexity increases and the chromatogram becomes populated with a larger number of peptides, as is the case for mAbs, problems with co-elutions will be evident. Method development for one biopharmaceutical product will not necessarily, and will probably not, be applicable to another product. These are the limitations to 1D LC that researchers encounter today.
We need a much larger peak capacity than the number of peptides in our digest. If we consider a digest with 100 peptides, a peak capacity of about 10,000 is required to have a good chance of separating all of them with chromatography. The introduction of high-end MS systems has alleviated the need for this extreme chromatographic performance but once methods are transferred from research to routine use, MS is out of the picture and the chromatography needs to do the job.
However, for relatively simple digests 1D LC can be suitable. More complex samples such as digests of very large biomolecules or mixtures thereof will seldom not require more than one analysis to completely profile the peptide map, even with MS installed. Combining different separations (that is, selectivities) is what 2D LC is essentially doing. This makes it a very valuable means to analyze such complex samples.
Q. What does two-dimensional chromatography offer that oneâdimensional chromatography cannot in the analytical study of large proteins?
A: The considerably increased separation power is what it is all about. Achieving resolution in the second dimension of compounds that are not separated in the first dimension is the most obvious advantage. However, 2D LC can be used in different modes.
First, there is comprehensive 2D LC (LC×LC) where the complete effluent from the first dimension is sampled in discrete fractions and each of these fractions is then analyzed in the second dimension. The main goal here is to increase peak capacity. In theory, the 2D LC peak capacity will be the product of each of the individual peak capacities.
This approach should increase separation power dramatically. In reality, however, the total peak capacity needs to be corrected for what is called undersampling and incomplete orthogonality. This has a serious impact and the practical peak capacity is significantly lower than the theoretical peak capacity. Nonetheless, the peak capacity in comprehensive 2D LC will be considerably higher than what can be achieved with 1D LC. This high peak capacity gives a comprehensive view of the sample constituents and resolves compounds that are not resolved with only the first- or second-dimension selectivity. The goal here could be to create a generic method to screen mAb digests for differences in amino acid sequence or post-translational modifications.
Other 2D LC techniques are based on heart-cutting approaches where only one or a number of fractions (peaks) are transferred to the second dimension. The advantage of such a method is that the second dimension analysis time, and therefore chromatographic performance, is more or less detached from the first dimension. In comprehensive LC×LC, the available time to perform the second dimension separation is very limited and, consequently, the full potential of this second dimension is unable to be exploited. This is not the case in heartcutting 2D LC approaches, so the choice of analytical conditions (column dimension, flow-rate, analysis time) can be tailored according to the analytical needs at hand. Applications for largeâbiomolecule separations include the combination of a first dimension ion-exchange mechanism with a secondâdimension MS-compatible reversed-phase separation. The inorganic salt present in the first dimension is removed after passing through the second dimension. In this way, a specific peak containing one or more peptides or proteins can be transferred on-line to the second dimension where an additional separation can take place before MS detection. Various ways to do this are available and the user-friendliness is superior compared to off-line methods where the peaks of interest have to be collected and then re-analyzed on another system following some manipulation.
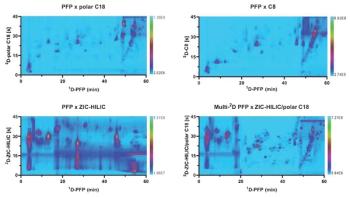
Q. Your research tested three different 2D LC combinations, namely strong cation exchange × reversedâphase LC, reversed-phase LC × reversed phase LC, and hydrophilic interaction liquid chromatography (HILIC) × reversed-phase LC. Did any of these combinations stand out as particularly effective for peptide mapping of large proteins?
A: Both reversed-phase LC × reversedâphase LC and strong cation exchange × reversedâphase LC are very suitable. In reversedâphase LC × reversedâphase LC, compatibility of both dimensions is excellent and efficiency on both columns is relatively high. It is the easiest of these three combinations and results in a robust method with high peak capacity. One of the disadvantages is that full orthogonality is impossible to achieve because of the similarity in separation mechanisms. However, there are plenty of tools to incorporate some orthogonality into this approach, by modifying factors such as the stationary phase, mobile phase pH and organic modifier type, and temperature. A simple option is to modify the second dimension gradient during the analysis. In a reversed-phase LC × reversed-phase LC set up this makes sense because compounds that are more strongly retained on the first dimension will also require stronger elution conditions in the second dimension. Combining all of these features results in powerful methods for detailed peptide mapping.
The strong cation exchange × reversedâphase LC combination will provide better orthogonality because the separation mechanisms are not correlated. Peptide and protein first dimension retention will be driven mainly by ionic interactions, while in the second dimension hydrophobicity of the compound will be the major factor for retention and separation. Strong cation exchange for peptides and proteins is generally performed using aqueous mobile phases, which makes the transfer to and injection onto the second dimension relatively straightforward. Possible drawbacks of this approach include the low chromatographic efficiency some strong cation exchange columns can provide. Also, the high buffer and salt load that is generally used in strong cation exchange separation of peptides and proteins will be introduced onto the second dimension and potentially the MS detector. The choice of a good first dimension column can obviously overcome the first drawback. The potential interference of inorganic mobile phase additives can somewhat be minimized by selecting the relevant window in which 2D LC needs to be performed and by using an additional diverter valve on the mass spectrometer.
HILIC × reversed-phase LC is the most difficult combination of the three. It will provide good orthogonality but there can be some compatibility issues. Since HILIC operates with high acetonitrile amounts, the transfer of these fractions onto the reversed-phase second dimension can cause polar peptides to breakthrough. However, various modifications can be made to reduce or even avoid this. It is definitely an interesting approach but I would advise this combination only in the case of specific analytical challenges such as differentiation of protein and peptide glycosylation. For general work, the reversed-phase LC × reversedâphase LC and strong cation exchange × reversedâphase LC combinations will be much easier to work with.
Q. You considered two options (formic acid and trifluoroacetic acid [TFA]) when replacing the mobile phase for MS detection. However, as noted, both options have potential drawbacks. Are there any alternatives that could have been used and, if so, in what circumstances would you recommend their use?
A: I would like to go back one step before answering this. For peptide mapping and protein analysis with reversed-phase LC, the use of a water–acetonitrile mobile phase with trifluoroacetic acid (TFA) is still frequently used. The use of TFA results in good retention for polar peptides because of its ion-pairing properties. However, it can produce interfering noise on the baseline when ultraviolet (UV) detection is used. This is a result of absorbance of UV light at 214 nm. Formic acid has a similar though less hindering effect. When running very fast gradients (generally around 30 s) as we do in comprehensive 2D LC, the excessive baseline noise and drift will lead to poor quality 2D LC plots. It will be difficult to detect small compounds in such data. This can be partly overcome by subtracting blank runs but this is limited and not very convenient. This is the reason we prefer to use phosphoric acid in the second dimension reversed-phase LC. With this additive a stable baseline is obtained and retention and selectivity are very close to those acquired with formic acid. In my opinion it is an excellent choice for 2D LC when a UV detector or diode-array detector (DAD) is used.
When MS is required, the use of phosphate or other inorganic mobile phase additives should be avoided. Here we need to replace phosphoric acid with a volatile and organic alternative. It is known that the ion-pairing effect of TFA can lead to ionization suppression. Formic acid is an excellent alternative because it will give nearly identical selectivity compared to phosphoric acid. So by using phosphate in UV work and formate in MS work we can easily compare results and identify peptides and proteins detected by UV based on their MS and MS–MS spectra.
For the best MS sensitivity, when analyzing peptides and proteins, positive ionization is preferred in combination with an acidic mobile phase. This is also the reason why we use high pH reversedâphase LC (ammonium bicarbonate pH 8.2) as the first dimension in our reversed-phase LC × reversedâphase LC setup. Fractions of this separation are then transferred to acidic conditions, which is ideal for positive ionization with electrospray.
Q. The method you developed showed a particular aptitude for identity, purity, and comparability assessments of biopharmaceuticals and biosimilars.1 Are there any limiting factors in the use of this method?
A: The main practical limitations of 2D LC lie in the compatibility of the dimensions and column availability. For the analysis of peptides and proteins a combination of reversed-phase LC × reversed-phase LC or strong cation exchange × reversedâphase LC will hardly ever disappoint. While compatibility in this case is not an issue, in other combinations the efforts required to overcome incompatibility between two dimensions (for example, flow splitting, mobile phase addition, combinations, trapping) could make the technique less accessible for routine use. That is why we generally start with reversedâphase LC × reversed-phase LC or strong cation exchange × reversed-phase LC combinations.
In some cases it can be problematic to find a column with suitable dimensions, in addition to the required packing chemistry and particle size. This is where column manufacturers could step in. For the first dimension column we like to work with a column internal diameter between 1 mm and 2.1 mm. This is done to keep the first dimension flow rate as low as possible to decrease volume loading onto the second dimension. The range of available stationary phases packed in narrow- and especially in micro-bore format is limited and frequently the ideal column will not be found in a catalogue. Custom ordering can be the solution but this takes time and can be costly.
Second-dimension columns to be used in heartcutting 2D LC approaches can be of any dimension and the choice here is plentiful. For comprehensive 2D LC applications this is not the case. The second dimension column should be able to provide good quality chromatography in as short an analysis time as possible (below 1 min), preferably with reasonably low flow rates and back pressure: Short columns packed with small particles or with superficially porous particles seem to be the obvious choice here. However, one has to be aware that one comprehensive 2D LC analysis will result in numerous injections (typically 50–150 modulations/run) of relatively large volumes (20–80 µL) onto this second dimension column. For this reason, the choice of a stable stationary phase with a slightly larger particle size, and thus end frits, could be justified. The small loss in efficiency will be largely compensated for by the improved robustness and column lifetime. In my opinion the ideal column for second dimension for biopharmaceuticals and protein digests is about 2 mm to 3 mm wide and 30 mm to 50 mm long packed with 3.5-µm particles. The use of a larger internal diameter increases loading capacity from the first dimension but requires high flow rates to keep the analysis time low enough (typically 3–5 mL/min). Such high flow rates lead to significantly high solvent consumption (several hundred mL/analysis), which is a disadvantage from an environmental as well as an economical point of view.
Of course, the price of 2D LC instrumentation and consumables and dedicated software is also something to keep in mind. The dedicated data analysis software for multidimensional chromatography is powerful and very useful, but it is not as developed as software developed for protein and peptide characterization and identification. Both software platforms often have to be used in combination, which can make data analysis more complicated.
Q.Are you planning to develop this methodology further and perhaps attempt to address some of those issues?
A: We have recently published an application note on strong cation exchange × reversed-phase LC analyses of E.coli tryptic digest and intact proteins.3 Strong cation exchange conditions were optimized compared to the published data on the mAb digests. With the optimized conditions we were able to generate a practical peak capacity of about 2250 in less than 4 h. This is outstanding and very useful for complex samples such as large biomolecule digests.
We have also recently used 2D LC in host cell protein (HCP) characterization and in determining the pharmacokinetic properties of antibody fragments. An interesting overview of the potential of 2D LC for the analysis of mAbs has recently been provided by our group.4
The use of heartcutting techniques has already proven very useful for hyphenating methods with inorganic mobile phases with MS after desalting on the second dimension separation. A typical application is identification of impurities and unknowns detected in the first-dimension separation. We are now also investigating the use of heartcutting for large biomolecules. Several separation principles commonly used for protein characterization (reversed-phase LC, strong cation exchange, size-exclusion chromatography [SEC], hydrophobic interaction chromatography [HIC]) can be used in the first dimension and well-defined fractions or peaks can be transferred on-line and analyzed using reversed-phase LC–DAD–MS.
Multi-dimensional LC is very powerful and flexible and new ideas will always surface as applications are developed. I am convinced that many applications will follow in the near future.
References
- G. Vanhoenacker, I. Vandenheede, F. David, P. Sandra, and K. Sandra, Analytical and Bioanalytical Chemistry407(1), 355–366 (2015).
- K. Sandra and P. Sandra, Advances in Biopharmaceutical Analysis28(s10)(2015).
- G. Vanhoenacker, K. Sandra, I. Vandenheede, F. David, and P. Sandra, Agilent Application Note 5991-5179EN (2014).
- K. Sandra and P. Sandra, Bioanalysis7(22), 2843–2847 (2015).
Gerd Vanhoenacker is the LC Product Manager at the Research Institute for Chromatography (RIC) in Kortrijk, Belgium. He studied Pharmaceutical Sciences at the Katholieke Universtiteit Leuven and obtained a Ph.D. degree in Pharmaceutical Sciences from the Ghent University in 2004.
He is (co-)author of over 40 scientific papers covering different areas of separation science. His expertise includes liquid chromatography (HPLC, UHPLC), liquid chromatography–mass spectrometry (LC–MS), supercritical fluid chromatography (SFC), capillary electrophoresis (CE), and sample preparation.
In recent years a significant part of his activities have included method development for 2D LC. He has practical experience with 2D LC for the analysis of a variety of samples.
Articles in this issue
over 10 years ago
Pattern Modulation Offers Alternative to Pulse Modulation in GC×GCover 10 years ago
Eastern Analytical Symposium Summaryover 10 years ago
Metabolomic Detection of Early-Stage Ovarian Cancerover 10 years ago
Reproducibility of Research — Do We Have a Problem Houston?over 10 years ago
Vol 12 No 1 The Column January 19, 2016 Europe and Asia PDFover 10 years ago
Vol 12 No 1 The Column January 19, 2016 North American PDF



