- Special Issues-07-01-2011
- Volume 9
- Issue 3
Imaging Mass Spectrometry: Current Performance and Upcoming Challenges
A critical look into the future of imaging mass spectrometry is made by addressing some of the fundamental and technical challenges that still need to be overcome.
Since its introduction in the late 1990s, matrix-assisted laser desorption–ionization (MALDI) imaging mass spectrometry (MS) technology has witnessed a phenomenal expansion. Initially introduced for the mapping of intact proteins from fresh frozen tissue sections, imaging MS is now routinely applied to a wide range of compounds including peptides, proteins, lipids, metabolites, and xenobiotics. Numerous compound-specific sample preparation protocols and analytical strategies have been developed, including tissue sectioning and handling, automated matrix deposition approaches, and the emergence of in-situ tissue chemistries such as enzymatic digestion to access the proteome locked in formalin-fixed paraffin embedded tissue biopsies. Instrumentation for imaging MS has also greatly evolved. Taking into account past progress and current capabilities, a critical look into the future of imaging MS needs to be made by addressing some of the fundamental and technical challenges that still need to be overcome for the technology to be fully accepted as a mainstream analytical tool.
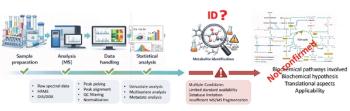
After a decade of development, matrix-assisted laser desorption–ionization (MALDI) imaging mass spectrometry (MS) is fast becoming a method of choice to map and characterize the molecular content of biological tissues (1–5). The first proof-of-principle experiments demonstrated the ability to spatially localize peptides and proteins from the direct analysis of thinly cut fresh frozen tissue sections (6,7). There is indeed a very unique correlation between the observed molecular signature pattern and the underlying histology. The principle of imaging MS is fairly simple and is explained in Figure 1. A thin section (~10 µm thick) is first cut from a frozen tissue block using a cryostat and thaw-mounted on a flat MALDI target plate. Serial sections are also cut and stained to align the imaging MS results with histological features. The next critical step is homogeneous matrix deposition directly on the section to be imaged. It has to be performed in a manner that minimizes delocalization of the analyte molecules of interest. To perform this process, several automated approaches can be used. After the matrix has dried the sample is analyzed by MALDI MS. Mass measurements on the section are performed in a systematic and automated manner. A grid array of fixed dimension and spatial resolution is defined and positioned over the section. At every grid coordinate, a mass spectrum is recorded by accumulating signals from multiple laser shots. For a chosen mass peak, an ion density map or image can then be extracted from the full data set by integrating and plotting its intensity in two dimensions as a function of spectrum coordinate. This ion image can be overlaid to the corresponding optical image generated from the matching stained serial section. From a single imaging MS measurement, several hundreds of different molecular images can be obtained.
Figure 1: Strategies for MALDI imaging MS sample preparation. See text for details.
Imaging MS has been used to study a wide range of biological systems. In particular, it has been used to study molecular expression and organization in diseases such as cancer (8–16) and neurologic disorders (17–20). Imaging MS also has been used extensively in developmental and fundamental biology (21–30). Another application area of growing interest is the mapping of xenobiotics and related metabolites in tissues (31–37). Not only can the exact location and absolute quantity of the drug be determined within a tissue specimen, but its effect on the proteome or lipidome can be monitored as a function of time or dose (38). There is also currently a growing interest for the monitoring of the metabolomic and proteomic content of plant tissues by imaging MS.
The State of The Art
Tissue Sample Collection and Processing
As for every MS experiment, the quality of the sample is of prime importance. Tissue specimens are ideally flash frozen immediately after sampling to minimize molecular degradation. It is also important to keep track of the orientation of the tissue specimen. Tissue samples can be loosely wrapped in aluminum foil to maintain the original shape as much as possible. The use of plastic tubes or tissue cassettes that are too small for the specimen should be avoided. After the samples are wrapped, flash freezing is typically performed by slow immersion (to avoid sample shattering) in liquid nitrogen or isopentane. Alternatively, samples can also be frozen on dry ice. Tissues should ideally be kept at –80 °C (or below) after freezing, until used for analysis (39,40).
Thin frozen sections are cut from the tissue blocks using a cryostat. Specimen cryostats are used for smaller specimens, whereas some applications call for full body cryostats such as whole-body imaging (35,41). The tissue blocks are maintained in place using polymeric embedding media and avoiding total immersion (when possible) to minimize contamination (39,40). Sections are typically cut with thicknesses of 8–20 µm. However, thinner sections are preferred to better visualize histological features. The sections are then thaw-mounted on the MALDI target plate. This is done by gently dragging the frozen section on the target plate previously cooled in the cryostat chamber. The section is then melted on the plate by warming its bottom surface on one's hand. To avoid any water condensation, this step is done within the cryostat chamber until all visible moisture has dried from the section. The plate can then be removed from the cryostat. The thaw-mounting step typically takes ~30 s (39).
Primarily developed for the imaging MS of proteins, the rinsing of tissue sections in organic solvents before matrix deposition has been shown to significantly improve signal quality and the lifetime of the sections (42,43). If lipid, metabolite, or drug imaging is the desired experiment, no washing should be performed. Several approaches ranging from "pipette" rinsing (42) to full immersion (43) have been demonstrated. Several organic solvent systems have been reported, but alcohol-based mixtures are most often employed (43). The rinsing of the sections has a dual effect. First, it eliminates physiological salts and a major part of the lipid component of the sections. This favors the desorption–ionization of proteins. Second, it stabilizes the protein content of the sections and allows storage for up to a few days (typically under desiccating conditions) without any major signal variations or loss (43).
Homogeneous Matrix Deposition
As stated above, matrix deposition on tissue sections needs to be performed without inducing any significant migration or delocalization of the analyte molecules investigated. The proper choices of matrix and solvent system have to be first considered as a function of the class of analyte molecules investigated and the MS instrumentation polarity used. Historically, the first coatings were performed by dragging a larger drop of matrix solution on the section in a cold-room environment. Although this approach can lead to some success (7), the delocalization of some proteins was high. Spray methods were also introduced early on, which allowed a better control of matrix deposition and thickness. Manual pneumatic spray (using an artist paintbrush or a thin-layer chromatography venturi flask) allows users to rapidly coat sections from small tissue specimens as well as from whole animals (35,41). It is still highly used today. Automated spray systems have emerged on the market (such as the TM sprayer from Leap Technologies, Carrboro, North Carolina) allowing a better control of spray flow, temperature, and deposition pattern (44). Another device uses a very fine aerosol of matrix to coat sections (ImagePrep system from Bruker Daltonics Inc., Billerica, Massachusetts). This mist redeposits on the section in a controlled environment and the matrix thickness is automatically controlled using a light-scattering device (45). With spray approaches for matrix deposition, the length of the matrix crystals forming on tissue are ≥10 µm.In this case, for most modern MALDI MS instruments the spatial resolution for imaging MS is therefore only limited by the diameter of the MALDI laser beam on target.
A second matrix deposition strategy developed of MALDI imaging MS is to print high density microdroplet arrays on the sections in an automated manner (Figures 1 and 2). By accumulation of several tens of picoliter volume droplets (~100 pL) at every array coordinate, high crystal density matrix microspots of <200-µm diameter are generated on the sections. Because matrix microspots do not touch, sample delocalization between two spots is eliminated. However, the center-to-center spacing between adjacent microspots defines the imaging MS resolution. Several different matrix printing systems have been developed. The first system operates with a piezo print head (CHiP 1000 chemical inkjet printer system from Shimadzu Corporation, Columbia, Maryland) (39,46,47). Matrix (initially contained in a small reservoir) is ejected on the sections mounted on a fast x,y stage system. At every grid coordinate, a picodroplet is collected. Printing stops after a predetermined number of print cycles over the array has been reached to build the microspots. The second system operates using acoustic energy to vertically eject picodroplets from a matrix reservoir (48). In this case, the section is mounted on a fast x,z stage and held inverted over the reservoir. In the same manner, droplets are collected at every grid coordinate (Figure 2). Even if resolution is limited with respect to matrix spray methods, the benefits are multiple. First, signal quality, particularly for proteins, is typically better in terms of signal-to-noise ratio and the overall number of peaks detected. Second, signals are extremely reproducible both in space and time. Printers are therefore ideal to obtain consistent and reproducible results when analyzing larger cohorts of samples that need to be investigated for longer periods of time. Other matrix printing systems, some homebuilt and others commercially available, have also been reported (49–51).
Figure 2: MALDI imaging MS of proteins from a coronal mouse brain section after automated droplet array matrix deposition using acoustic-based ejection (matrix: sinapinic acid, 275 µm center-to-center). Mass spectra are acquired from every microspots by TOF-MS and ion images are reconstructed when plotting peak intensities at specific m/z of interest as a function of grid array coordinates. Areas of signal expression can directly be correlated with histological features present in the section.
Two new dry on-tissue matrix deposition approaches have recently been proposed; however, these have proven to be extremely powerful for only the imaging MS of low-molecular-weight molecules, particularly phospholipids (52,53). Imaging MS of some metabolites and drug compounds have also been reported (54). The first approach consists in sublimating matrix on the tissue sections (Figure 3). This was first reported using a small sublimation apparatus coupled to a rough pumping system and a heating sand block (52). This allows for depositing an extremely homogeneous micrometer-thick layer of matrix on the section and the imaging MS of phospholipids with spatial resolutions in the low micrometer range (55). Other more elaborate setups and strategies have also been proposed (56,57). The second approach uses fine mesh sieves (~20 µm) to deposit very finely ground matrix on the tissue section (53). When the sieve is shaken, matrix passes through the mesh and deposits itself on the section. Surprisingly, a thin matrix layer forms that sticks to the section. In this case, coverage is usually not complete but allows imaging with a spatial resolution of tens of micrometers. Sieves of different diameter and mesh size are available and the shaking of the sieves can be automated.
Figure 3: MALDI imaging MS of phospholipids from a saggital mouse brain section after matrix deposition by sublimation (matrix: 2,5-dihydroxy cinnamic acid). Mass spectra are acquired by high-resolution TOF over the surface of the section with a spatial resolution of 100 µm. Shown are an H&E stained serial section, the section after the imaging process, and eight ion images obtained for different phospholipid species located in different brain substructures.
MALDI Imaging MS Instrumentation and Strategies
In the past decade, instrumentation for imaging MS has also greatly evolved. The first MS images were collected with MALDI time-of-flight (TOF) instruments capable of laser repetition rates of a few hertz (7). Today, modern TOF systems are capable of acquiring data in the kilohertz range with improved sensitivity, mass resolving power, and accuracy, significantly reducing acquisition time and improving image quality (45,58). Beyond TOF analyzers, other MALDI-based instruments have been used such as ion traps (59–61), quadrupole quadrupole TOFs (QqTOFs) (41), and trap-TOF (48). Ion mobility technology has also been used in conjunction with imaging MS (62,63). More recently, MALDI Fourier-transform ion cyclotron resonance (FT-ICR) and orbital trap mass spectrometers have been demonstrated to be valuable instruments for imaging MS at very high mass resolving power. These non-TOF based systems have proven to be extremely powerful for the imaging of lower-molecular-weight compounds such as lipids, drugs, and metabolites (17,31,64).
Compound imaging is generally performed in the "MS mode" where ion images are reconstructed based on the intensity of the corresponding MS signals (2,7). This approach is typically used in nontargeted or exploratory experiments in which the amount of information to be recovered from a given tissue sample is maximized. For example, with TOF systems protein imaging in the MS mode typically allows users to map from several hundred to, in some cases, as many as 1000 m/z peaks over two orders of magnitude in intensity (65). Lower-molecular-weight imaging with high resolving power instruments in the m/z range below 1000 allows users to map several thousands of compounds with signal intensities over three orders of magnitude (31).
Imaging MS also can be performed in a targeted approach using a single reaction monitoring (SRM) strategy (33). In this case, ion images are reconstructed based on the intensity of a predefined fragment ion from a targeted compound of interest. Imaging MS with a SRM approach introduces a high degree of specificity and sensitivity. For SRM imaging approaches, QqTOF, ion-trap, and triple-quadrupole MALDI MS systems are typically employed (36,37,41). For example, SRM imaging is often used to monitor the location and abundance of administered drug compounds across tissues, organs, and whole-body sections (35,41). Targeted imaging MS can also be performed using systems with high mass resolving power in which, in this case, images are reconstructed using the exact mass of the compounds (31).
MALDI Imaging MS Data Acquisition, Visualization, and Analysis
The critical aspects of MALDI imaging MS are automated data acquisition and image reconstruction and visualization. The first software for imaging MS was written in-house (66,67). This software allowed precise movement of the sample plate in TOF-MS instruments, automated data acquisition at every coordinate of a user-predefined grid array, and generated ion images for a preselected set of m/z values. Today, most manufacturers of MALDI MS instruments have proposed software solutions for imaging MS. Several MS data preprocessing tools have also been incorporated in the image acquisition and visualization software, including background removal, denoising, smoothing, and data normalization (typically by total ion current) subroutines (44). Postacquisition statistical software packages are also starting to be available that perform class comparisons between data sets extracted from different regions of interests within the tissue sections (68–70).
Current and Future Challenges
Although an extraordinary amount of progress has been made and large quantities of collective knowledge have been gained in the past decade, imaging MS is still not fully recognized as a mainstream analytical tool. To achieve this status, several key technical hurdles have to be overcome.
Imaging MS with High Specificity and Sensitivity
Most often, MALDI imaging MS is performed in a nontargeted manner and the more abundant biocompounds are primarily seen. For example, when imaging proteins by TOF-MS, the bulk of the information is seen in the molecular weight range below 20,000 where molecules are better detected and mass resolved. Further, with standard matrix preparation methods in a high organic content, most of the proteins seen are soluble and for the most part are known to be abundant in cells.
Methods for specifically targeting a given family of proteins need to be developed. A high-specificity imaging MS approach has been introduced in which protein recognition is done using a specific antibody previously coupled to a low-molecular-weight reporter mass tag via a photocleavable linker molecule (71–73). After reaction, imaging MS is performed by releasing the reporter mass tag under laser desorption ionization conditions. An image is reconstructed for the mass tag, which acts as a surrogate marker of the protein. Beyond on-tissue section matrix deposition, the automated liquid dispensing devices described above can be used to do in-situ chemistries. This approach has been used extensively to perform in-situ enzymatic digestions before imaging MS. Trypsin can be first homogeneously deposited on the sections, which causes proteins to be locally digested into peptides (74). After matrix deposition, proteins are imaged via the localization of their corresponding peptides; in particular, this technique has allowed the imaging MS of higher-molecular-weight proteins. This approach has also opened the way for the proteomic analysis of formalin fixed paraffin embedded tissue sections by MALDI imaging MS (49,75).
Although some restrictions in terms of reaction temperature and pressure exist with current liquid handling instrumentation, the possibilities for in-situ chemistries applied to proteins and other endogenous biocompounds to gain in specificity and sensitivity are numerous. Chemistry also can be introduced on the target plate on which sections are mounted. Early publications on MALDI imaging MS introduced this concept in which tissues were blotted on different surfaces (C18 coated, polyethylene, or PVDF) and the imprints were profiled or imaged (6,76–78). The specific capture of biocompounds using functionalized surfaces is therefore possible. Here, again, capture chemistry design is endless, from very generic to very specific.
Matrix Deposition and MS Instrumentation
MS instrumentation is continually improving with enhanced resolving power, sensitivity, and mass accuracy. Beyond enhanced spectral quality, two important factors have to be considered for MALDI imaging MS, namely laser spot size on target and speed of acquisition. One of the goals of imaging MS is to be able to image at cellular-length scales (for example, ≤10 µm of spatial resolution). Several homebuilt systems have shown that MALDI imaging MS at or below 5 µm is possible (55,67,79). Today, state of the art commercially available MALDI-TOF-MS systems are capable of offering laser beam size on target in the range of 20 µm in diameter (45). It is therefore foreseeable that the next generation of instruments should be capable of imaging in the single-cell regime.
To image at cellular resolution implies that we have the ability to deposit matrix in a homogeneous manner without lateral analyte migration beyond a few micrometers. Only matrix sublimation has shown such potential; unfortunately it is not currently applicable to protein imaging directly. Ideally, matrix deposition must be performed to locally form a homogeneous monolayer of consistent micrometer size crystals covering close to 100% of the surface of the section, which upon laser irradiation generate strong protein ion signals. Clearly current matrix deposition methodologies cannot achieve this coverage and significant efforts need to be made in sample preparation. Smaller irradiated surfaces mean that less material is ablated and fewer ions are formed, so sensitivity may become an issue especially in the higher-molecular-weight range. Second, to investigate a given area at higher spatial resolution implies that the number of pixels per imaging MS experiment will significantly increase. However, total imaging time still must stay manageable so imaging speed becomes a critical parameter. In this regard, modern TOF-MS systems show the highest imaging speed performance by offering laser repetition rates in the kilohertz range, which greatly limits total data acquisition time (45,58).
Processing and Statistical Analysis of Imaging MS Data
In most instances, when considering the amount of information published from imaging MS data sets, only a handful of images from various compounds are typically used. The vast majority of the information contained in the data sets is untapped. Imaging MS data sets can contain anywhere from a few hundred to a few thousand different ion images and manual extraction becomes virtually impossible. Automated approaches to extract ion images need to be incorporated in the next generation of imaging software (80). Furthermore, these sets of images need to be organized or classified based on alignment with histological features or predefined regions of interests from the tissue sections.
Some statistical approaches already exist in which classification based on principal component analysis and other approaches is possible (69,70,81,82). Even after imaging MS, compound identification (see below), and correlation with histology, results may not be initially informative with respect to the biology of the sample. This analysis is especially difficult to do when lipid and protein imaging MS data sets acquired from the same tissue samples are compared. It is necessary to integrate these results using a systems biology approach to interpret and understand the studied biological systems.
Compound Identification and Quantitation
Compound identification is currently the biggest bottleneck of imaging MS. Whereas within the course of a day hundreds of m/z-specific images can be generated, it may take weeks to identify and validate the identity of the corresponding biomolecules. For lower-molecular-weight compounds, a first-pass identification is generally possible by MALDI MS-MS directly from the tissue section. This has been nicely demonstrated for some metabolites, lipids, and peptides (74,83,84). However, this approach is applicable particularly if the precursor mass signal is strong and reasonably easy to preselect. It becomes more challenging to identify compounds coming from lower intensity signals especially in the presence of stronger intensity isobaric ions. This is typically the case when imaging lipids, where within a single isobar several species can be found with very different signal intensities.
Protein identification poses a bigger challenge because a MALDI MS-MS approach cannot be used. In most instances, a classical top-down protein identification approach must be applied. Proteins are first extracted from the tissue sample and separated by chromatography. After protein isolation and enzymatic (trypsin) digestion, the corresponding peptides are mapped and sequenced by MS and the results used to identify the corresponding protein (23,39,77). Such strategies are robust, but time consuming. Solutions to streamline identification have been proposed, such as incorporating modern protein fragmentation methods after the initial separation. There is, however, room for improvement. Validation of the correct identification is also a critical step. For imaging MS, validation is typically done by immunohistochemistry (IHC) (23). In this case, the imaging MS results are compared to the imaging results after IHC. Modern alternative validation strategies by MALDI SRM following a set of target peptides from the identified protein are also possible.
Ion images from a given compound already give some information about its distribution and relative abundance across a tissue section. Approaches for the absolute quantitation of compounds directly from tissue sections also need to be developed. Quantitation has already proven possible by MALDI imaging MS for lower-molecular-weight compounds, namely administered pharmaceuticals. In this case, signal intensity was found directly proportional to the local drug concentration (37,85). Absolute quantitation should also be possible by the addition of internal standards. Absolute protein quantitation strategies using a MALDI SRM approach after addition of an isotopically labeled standard peptide need to be developed.
Conclusions
Since its introduction in the late 1990s, collective efforts from a growing numbers of laboratories worldwide in collaboration with MS instrumentation manufacturers have allowed the development of robust methods for the imaging by MALDI MS of biocompounds present in tissue specimens. These include tissue specimen handling and sectioning, homogeneous matrix deposition, automated data acquisition, image reconstruction and visualization, and data analysis.
The growing interest in the technology clearly illustrates the potential of MALDI imaging MS to become a mainstream analytical tool in bioscience research. To achieve this, new or improved methodologies still need to be developed for high-throughput imaging MS at cellular-length scales. New bioinformatic tools still need to be developed to quarry and understand the biological meaning of the hundreds of molecular images acquired from tissue samples. Improved rapid and sensitive strategies for molecular identification and quantitation need to be developed.
The past decade has been important for the advent of MALDI imaging MS. With a growing interest from the scientific community, the next decade will be rich in development and will constantly push back the limits of the technology.
Acknowledgments
All of the data presented in this article were acquired at Vanderbilt University (Nashville, Tennessee). The author would like to thank professor Richard Caprioli and his research group for more than a decade of collaborative efforts and the production of outstanding research.
References
(1) D.S. Cornett, M.L. Reyzer, P. Chaurand, and R.M. Caprioli, Nat. Methods 4, 828–833 (2007).
(2) L.A. McDonnell and R.M.A. Heeren, Mass Spectrom. Rev. 26, 606–643 (2007).
(3) E.H. Seeley and R.M. Caprioli, Proc. Natl. Acad. Sci. U.S.A. 105, 18126–18131 (2008).
(4) E.R.A. van Hove, D.F. Smith, and R.M.A. Heeren, J. Chromato. A 1217, 3946–3954 (2010).
(5) E.H. Seeley and R.M. Caprioli, Trends Biotechnol. 29, 136–143 (2011).
(6) R.M. Caprioli, T.B. Farmer, and J. Gile, Anal. Chem. 69, 4751–4760 (1997).
(7) M. Stoeckli, P. Chaurand, D.E. Hallahan, and R.M. Caprioli, Nat. Med. 7, 493–496 (2001).
(8) R.M. Caprioli, Proteomics 8, 3679–3680 (2008).
(9) L.H. Cazares, D. Troyer, S. Mendrinos, R.A. Lance, J.O. Nyalwidhe, H.A. Beydoun, M.A. Clements, R.R. Drake, and O.J. Semmes, Clin. Cancer Res. 15, 5541–5551 (2009).
(10) P. Chaurand, M.E. Sanders, R.A. Jensen, and R.M. Caprioli, Am. J. Pathol. 165, 1057–1068 (2004).
(11) P. Chaurand, S.A. Schwartz, and R.M. Caprioli, J. Proteome Res. 3, 245–252 (2004).
(12) K. Chughtai and R.M.A. Heeren, Chem. Rev. 110, 3237–3277 (2010).
(13) R. Lemaire, S.A. Menguellet, J. Stauber, V. Marchaudon, J.P. Lucot, P. Collinet, M.O. Farine, D. Vinatier, R. Day, P. Ducoroy, M. Salzet, and I. Fournier, J. Proteome Res. 6, 4127–4134 (2007).
(14) L.A. McDonnell, G.L. Corthals, S.M. Willems, A. van Remoortere, R.J.M. van Zeijl, and A.M. Deelder, J. Proteomics 73, 1921–1944 (2010).
(15) Y. Morita, K. Ikegami, N. Goto-Inoue, T. Hayasaka, N. Zaima, H. Tanaka, T. Uehara, T. Setoguchi, T. Sakaguchi, H. Igarashi, H. Sugimura, M. Setou, and H. Konno, Cancer Sci. 267–273 (2010).
(16) K. Schwamborn and R.M. Caprioli, Mol. Oncol. 4, 529–538 (2010).
(17) R.R. Landgraf, M.C.P. Conaway, T.J. Garrett, P.W. Stacpoole, and R.A. Yost, Anal. Chem. 81, 8488–8495 (2009).
(18) J. Pierson, J.L. Norris, H.R. Aerni, P. Svenningsson, R.M. Caprioli, and P.E. Andren, J. Proteome Res. 3, 289–295 (2004).
(19) K. Skold, M. Svensson, A. Nilsson, X. Zhang, K. Nydahl, R.M. Caprioli, P. Svenningsson, and P.E. Andren, J. Proteome Res. 5, 262–269 (2006).
(20) M. Wisztorski, D. Croix, E. Macagno, I. Fournier, and M. Salzet, Dev. Neurobiol. 68, 845–858 (2008).
(21) K.E. Burnum, D.S. Cornett, S.M. Puolitaival, S.B. Milne, D.S. Myers, S. Tranguch, H.A. Brown, S.K. Dey, and R.M. Caprioli, J. Lipid Res. 50, 2290–2298 (2009).
(22) K.E. Burnum, S. Tranguch, D. Mi, T. Daikoku, S.K. Dey, and R.M. Caprioli, Endocrinology 149, 3274–3278 (2008).
(23) P. Chaurand, S. Fouchecourt, B.B. DaGue, B.J. Xu, M.L. Reyzer, M.C. Orgebin-Crist, and R.M. Caprioli, Proteomics 3, 2221–2239 (2003).
(24) P. Chaurand, M.A. Rahman, T. Hunt, J.A. Mobley, G. Gu, J.C. Latham, R.M. Caprioli, and S. Kasper, Mol. Cell. Proteomics 7, 411–423 (2008).
(25) R.B. Chen, L.M. Hui, R.M. Sturm, and L.J. Li, J. Am. Soc. Mass Spectrom. 20, 1068–1077 (2009).
(26) A.C. Grey, P. Chaurand, R.M. Caprioli, and K.L. Schey, J. Proteome Res. 8, 3278–3283 (2009).
(27) A.C. Grey, A.K. Gelasco, J. Section, R.A. Moreno-Rodriguez, E.L. Krug, and K.L. Schey, Anat. Rec. A. Discov. Mol. Cell. Evol. Biol. 293, 821–828 (2010).
(28) J. Han and K.L. Schey, Invest. Ophthalmol. Vis. Sci. 47, 2990–2996 (2006).
(29) K.D. Herring, S.R. Oppenheimer, and R.M. Caprioli, Semin. Nephrol. 27, 597–608 (2007).
(30) L. Minerva, S. Clerens, G. Baggerman, and L. Arckens, Proteomics 8, 3763–3774 (2008).
(31) D.S. Cornett, S.L. Frappier, and R.M. Caprioli, Anal. Chem. 80, 5648–5653 (2008).
(32) Y. Hsieh, J. Chen, and W.A. Korfmacher, J. Pharmacol. Toxicol. Methods 55, 193–200 (2007).
(33) M.L. Reyzer and R.M. Caprioli, Curr. Opin. Chem. Biol. 11, 29–35 (2007).
(34) M. Rudin, M. Rausch, and M. Stoeckli, Mol. Imaging Biol. 7, 5–13 (2005).
(35) M. Stoeckli, D. Staab, A. Schweitzer, J. Gardiner, and D. Seebach, J. Am. Soc. Mass Spectrom. 18, 1921–1924 (2007).
(36) Y. Sugiura and M. Setou, J. Neuroimmune Pharmacol. 5, 31–43 (2010).
(37) B. Prideaux, V. Dartois, D. Staab, D.M. Weiner, A. Goh, L.E. Via, C.E. Barry, and M. Stoeckli, Anal. Chem. 83, 2112–2118 (2011).
(38) M.L. Reyzer, R.L. Caldwell, T.C. Dugger, J.T. Forbes, C.A. Ritter, M. Guix, C.L. Arteaga, and R.M. Caprioli, Cancer Res. 64, (2004).
(39) P. Chaurand, J.L. Norris, D.S. Cornett, J.A. Mobley, and R.M. Caprioli, J. Proteome Res. 5, 2889–2900 (2006).
(40) S.A. Schwartz, M.L. Reyzer, and R.M. Caprioli, J. Mass Spectrom. 38, 699–708 (2003).
(41) S. Khatib-Shahidi, M. Andersson, J.L. Herman, T.A. Gillespie, and R.M. Caprioli, Anal. Chem. 78, 6448–6456 (2006).
(42) R. Lemaire, M. Wisztorski, A. Desmons, J.C. Tabet, R. Day, M. Salzet, and I. Fournier, Anal. Chem. 78, 7145–7153 (2006).
(43) E.H. Seeley, S.R. Oppenheimer, D. Mi, P. Chaurand, and R.M. Caprioli, J. Am. Soc. Mass Spectrom. 19, 1069–1077 (2008).
(44) J.L. Norris, D.S. Cornett, J.A. Mobley, M. Andersson, E.H. Seeley, P. Chaurand, and R.M. Caprioli, Int. J. Mass Spectrom. 260, 212–221 (2007).
(45) M. Lagarrigue, M. Becker, R. Lavigne, S.-O. Deininger, A. Walch, F. Aubry, D. Suckau, and C. Pineau, Mol. Cell. Proteomics 10, 1–11 (2011).
(46) H.R. Aerni, D.S. Cornett, and R.M. Caprioli, Anal. Chem. 78, 827–834 (2006).
(47) D.S. Cornett, J.A. Mobley, E.C. Dias, M. Andersson, C.L. Arteaga, M.E. Sanders, and R.M. Caprioli, Mol. Cell. Proteomics 5, 1975–1983 (2006).
(48) S. Shimma, M. Furuta, K. Ichimura, Y. Yoshida, and M. Setou, Surf. Interface Anal. 38, 1712–1714 (2006).
(49) R. Lemaire, A. Desmons, J.C. Tabet, R. Day, M. Salzet, and I. Fournier, J. Proteome Res. 6, 1295–1305 (2007).
(50) Y.F. Chen, J. Allegood, Y. Liu, E. Wang, B. Cachon-Gonzalez, T.M. Cox, A.H. Merrill, and M.C. Sullards, Anal. Chem. 80, 2780–2788 (2008).
(51) A. Vegvari, T.E. Fehniger, L. Gustavsson, A. Nilsson, P.E. Andren, K. Kenne, J. Nilsson, T. Laurell, and G. Marko-Varga, J. Proteomics 73, 1270–1278 (2010).
(52) J.A. Hankin, R.M. Barkley, and R.C. Murphy, J. Am. Soc. Mass Spectrom. 18, 1646–1652 (2007).
(53) S.M. Puolitaival, K.E. Burnum, D.S. Cornett, and R.M. Caprioli, J. Am. Soc. Mass Spectrom. 19, 882–886 (2008).
(54) R.J.A. Goodwin, P. Scullion, L. MacIntyre, D.G. Watson, and A.R. Pitt, Anal. Chem. 82, 3868–3873 (2010).
(55) P. Chaurand, D.S. Cornett, P.M. Angel, and R.M. Caprioli, Mol. Cell. Proteomics 10, 10.1074/mcp.O1110.004259, 004251-004211 (2011).
(56) L.J.M. Dekker, J.J.A. van Kampen, M.L. Reedijk, P.C. Burgers, R.A. Gruters, A. Osterhaus, and T.M. Luider, Rapid Commun. Mass Spectrom. 23, 1183–1188 (2009).
(57) W. Bouschen, O. Schulz, D. Eikel, and B. Spengler, Rapid Commun. Mass Spectrom. 24, 355–364 (2010).
(58) J.M. Spraggins and R.A. Caprioli, J. Am. Soc. Mass Spectrom. 22, 1022–1031 (2011).
(59) P.D. Verhaert, M.C.P. Conaway, T.M. Pekar, and K. Miller, Int. J. Mass Spectrom. 260, 177–184 (2007).
(60) T.J. Garrett, M.C. Prieto-Conaway, V. Kovtoun, H. Bui, N. Izgarian, G. Stafford, and R.A. Yost, Int. J. Mass Spectrom. 260, 166–176 (2007).
(61) D.M. Drexler, T.J. Garrett, J.L. Cantone, R.W. Diters, J.G. Mitroka, M.C. Prieto Conaway, S.P. Adams, R.A. Yost, and M. Sanders, J. Pharmacol. Toxicol. Methods 55, 279–288 (2007).
(62) S.N. Jackson, M. Ugarov, T. Egan, J.D. Post, D. Langlais, J.A. Schultz, and A.S. Woods, J. Mass Spectrom. 42, 1093–1098 (2007).
(63) J. Stauber, L. MacAleese, J. Franck, E. Claude, M. Snel, B.K. Kaletas, I. Wiel, M. Wisztorski, I. Fournier, and R.M.A. Heeren, J. Am. Soc. Mass Spectrom. 21, 338–347 (2010).
(64) I.M. Taban, A.F.M. Altelaar, Y.E.M. Van der Burgt, L.A. McDonnell, R.M.A. Heeren, J. Fuchser, and G. Baykut, J. Am. Soc. Mass Spectrom. 18, 145–151 (2007).
(65) P. Chaurand and R.M. Caprioli, Electrophoresis 23, 3125–3135 (2002).
(66) M. Stoeckli, T.B. Farmer, and R.M. Caprioli, J. Am. Soc. Mass Spectrom. 10, 67–71 (1999).
(67) B. Spengler and M. Hubert, J. Am. Soc. Mass Spectrom. 13, 735–748 (2002).
(68) K. Schwamborn, R.C. Krieg, M. Reska, G. Jakse, R. Knuechel, and A. Wellmann, Int. J. Mol. Med. 20, 155–159 (2007).
(69) S.O. Deininger, M.P. Ebert, A. Futterer, M. Gerhard, and C. Rocken, J. Proteome Res. 7, 5230–5236 (2008).
(70) T. Alexandrov, M. Becker, S.O. Deininger, G. Ernst, L. Wehder, M. Grasmair, F. von Eggeling, H. Thiele, and P. Maass, J. Proteome Res. 9, 6535–6546 (2010).
(71) R. Lemaire, J. Stauber, M. Wisztorski, C. Van Camp, A. Desmons, M. Deschamps, G. Proess, I. Rudlof, A.S. Woods, R. Day, M. Salzet, and I. Fournier, J. Proteome Res. 6, 2057–2067 (2007).
(72) G. Thiery, E. Anselmi, A. Audebourg, E. Darii, M. Abarbri, B. Terris, J.C. Tabet, and I.G. Gut, Proteomics 8, 3725–3734 (2008).
(73) G. Thiery, M.S. Shchepinov, E.M. Southern, A. Audebourg, V. Audard, B. Terris, and L.G. Gut, Rapid Commun. Mass Spectrom. 21, 823–829 (2007).
(74) M.R. Groseclose, M. Andersson, W.M. Hardesty, and R.M. Caprioli, J. Mass Spectrom. 42, 254–262 (2007).
(75) M.R. Groseclose, P. Chaurand, P.P. Massion, and R.A. Caprioli, Proteomics 8, 3715 - 3724 (2008).
(76) P. Chaurand, M. Stoeckli, and R.M. Caprioli, Anal. Chem. 71, 5263–5270 (1999).
(77) P. Chaurand, B.B. DaGue, R.S. Pearsall, D.W. Threadgill, and R.M. Caprioli, Proteomics 1, 1320–1326 (2001).
(78) N. Goto-Inoue, T. Hayasaka, Y. Sugiura, T. Taki, Y.T. Li, M. Matsumoto, and M. Setou, J. Chromato. B-Anal. Technol. Biomed. Life Sci. 870, 74–83 (2008).
(79) P. Chaurand, K.E. Schriver, and R.M. Caprioli, J. Mass Spectrom. 42, 476–489 (2007).
(80) L.A. McDonnell, A. van Remoortere, N. de Velde, R.J.M. van Zeijl, and A.M. Deelder, J. Am. Soc. Mass Spectrom. 21, 1969–1978 (2010).
(81) G. McCombie, D. Staab, M. Stoeckli, and R. Knochenmuss, Anal. Chem. 77, 6118–6124 (2005).
(82) F.Q. Zhang and D. Hong, Stat. Med. 30, 753–768 (2011).
(83) F. Benabdellah, D. Touboul, A. Brunelle, and O. Laprevote, Anal. Chem. 81, 5557–5560 (2009).
(84) S.N. Jackson, H.Y.J. Wang, and A.S. Woods, J. Am. Soc. Mass Spectrom. 16, 2052–2056 (2005).
(85) M.L. Reyzer, Y. Hsieh, K. Ng, W.A. Korfmacher, and R.M. Caprioli, J. Mass Spectrom. 38, 1081–1092 (2003).
Pierre Chaurand is with the University of Montreal, Montreal, Canada.
Articles in this issue
almost 15 years ago
High-Definition Screening for Boar Taint in Fatback Samples Using GC–MSalmost 15 years ago
Mass Spectrometry in Analytical Lipidomics